It seems that biologists are great at creating -omes. There’s the genome, the complete set of genetic information held in a plant’s DNA. There’s the transcriptome, the sum of all the RNA expressed by the genome, that can tell you which genes are active. Some biologists are now studying the ionome, but what is the ionome?
The ionome is defined as the elemental composition of a subcellular structure, cell, tissue, organ or organism. It includes all elements (except hydrogen, H, and oxygen, O), whether essential or non-essential to life, in whatever chemical form they occur. Ionomics is the study of the ionome.
Biologists have studied the effects of genotype and environment on the elemental composition of plants for many years. What makes ionomics special is that it’s the study of many elements simultaneously, rather than just one element at a time. Every plant doesn’t just need nutrients, they need those nutrients in a specific balance. If they cannot balance their nutrients, plants perform less well and can even die.
So, an important factor in your garden is to plant plants that can balance the nutrients in the soil, and apply any fertilisers carefully. For example, phosphorus is usually considered a good nutrient for a plant, but have too much and it can impair a plant’s ability to take up other nutrients, such as zinc or manganese, that it also needs from the soil.
Since the ionome includes more than ions, it is actually a misnomer and some people prefer the term “elementome”. Plant nutritionists sometimes refer to the “functional ionome”, meaning the 14 elements (in addition to carbon, C, H and O) required by plants: nitrogen (N), phosphorus (P), potassium (K), calcium (Ca), magnesium (Mg), sulphur (S), chlorine (Cl), boron (B), iron (Fe), manganese (Mn), copper (Cu), zinc (Zn), nickel (Ni) and molybdenum (Mo). This is also referred to as the “nutriome” by some authors. Another sub-division of the ionome is the “metallome”, which comprises “metal and metalloid species”. This term has its own journal, Metallomics.
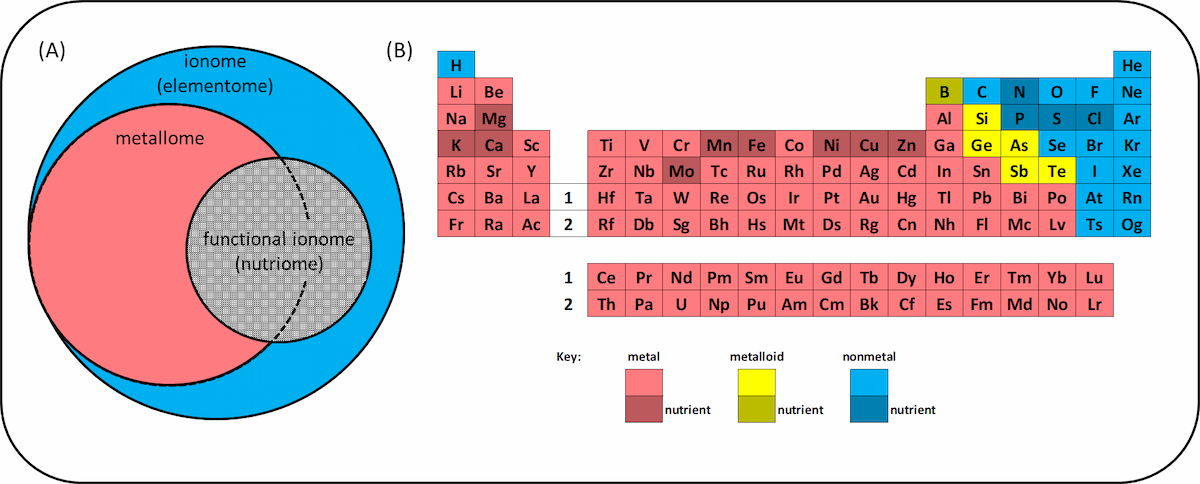
There are people working on the same problem, but using different terms. Does this matter if we know the words all mean broadly the same thing? I decided to search through the literature to find out.
On 5th July, 2021, I searched the Web of Science Core Collection with the terms “(ionome or elementome or nutriome or metallome or ionomics or elementomics or nutriomics or metallomics) and (plant or root or stem or leaf or leaves or shoot or fruit or seed or grain)”. This returned 380 articles (excluding corrections) on plants. Most of these were original articles. About 15% were reviews or editorial material.
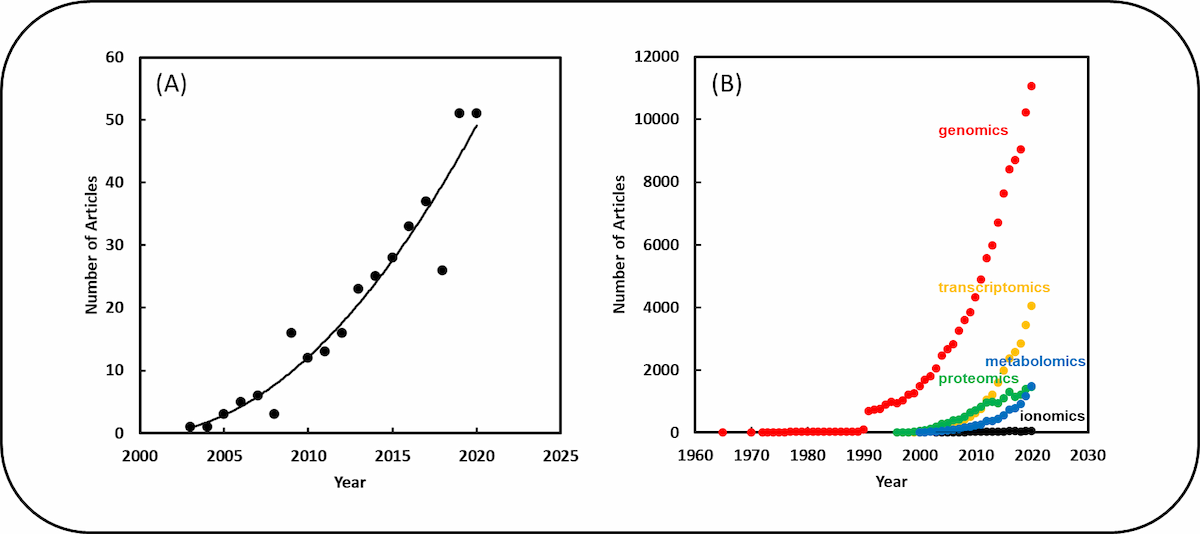
The most popular term was “ionome / ionomics”, which occurred in 326 articles, followed by “metallome / metallomics”, which occurred in 52 articles. The terms “elementome / elementomics” and “nutriome / nutriomics” occurred in only in 3 and 4 articles, respectively. The use of these terms has grown in popularity (Fig. 2) since the first use of the word “ionome” in an article by Kendal Hirschi (2003), but their use is clearly much less than the use of “genome / genomics”, “transcriptome / transcriptomics”, “proteome / proteomics” or “metabolome / metabolomics”, which have a longer history (Fig. 2).
David Salt (University of Nottingham) authored 27 of the articles retrieved by the search and his collaborators Ivan Baxter (Donald Danforth Plant Science Centre) and Bret Lahner (Purdue University) authored 23 and 11 articles, respectively. Other authors who contributed to a significant number of articles include Marco Aurelio Zezzi Arruda (Universidade Estadual de Campinas, 13 articles), Toshihiro Watanabe (Hokkaido University, 12 articles), Takuro Shinano (Hokkaido University, 10 articles) and Philip White (James Hutton Institute, 10 articles). The most active countries publishing on “ionomics” are the USA and China, which might simply reflect the greater investment of these countries in plant science.
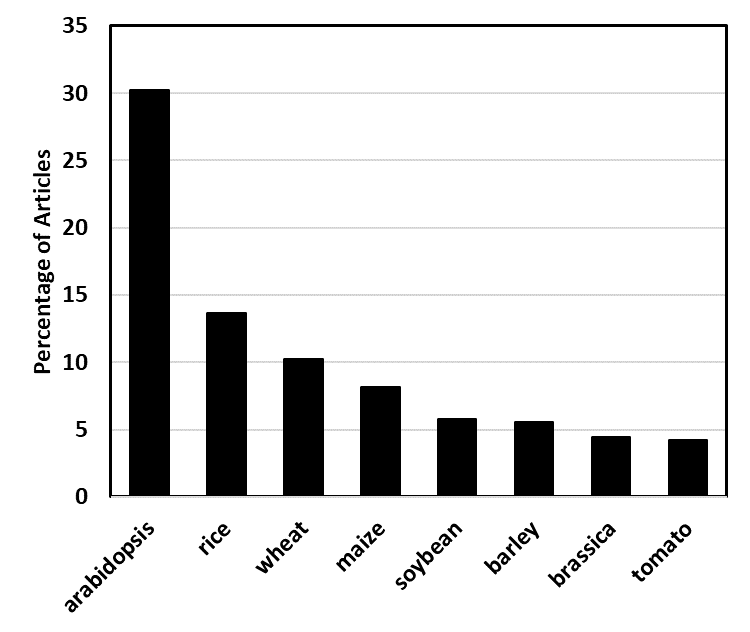
About half the number of articles retrieved covered the area of “Plant Sciences”, 17% were in “Agriculture”, 15% in “Ecology” and 12% in “Molecular Biology”. Almost one third of the articles described the ionomics of arabidopsis, whilst another third focussed on the cereals, rice, wheat, maize and barley (Fig. 3). Other well-studied plants include brassicas, soybean and tomato. Interestingly, a greater than expected number of papers from the USA focussed on arabidopsis and a greater than expected number of papers from China focussed on cereals, rice in particular.
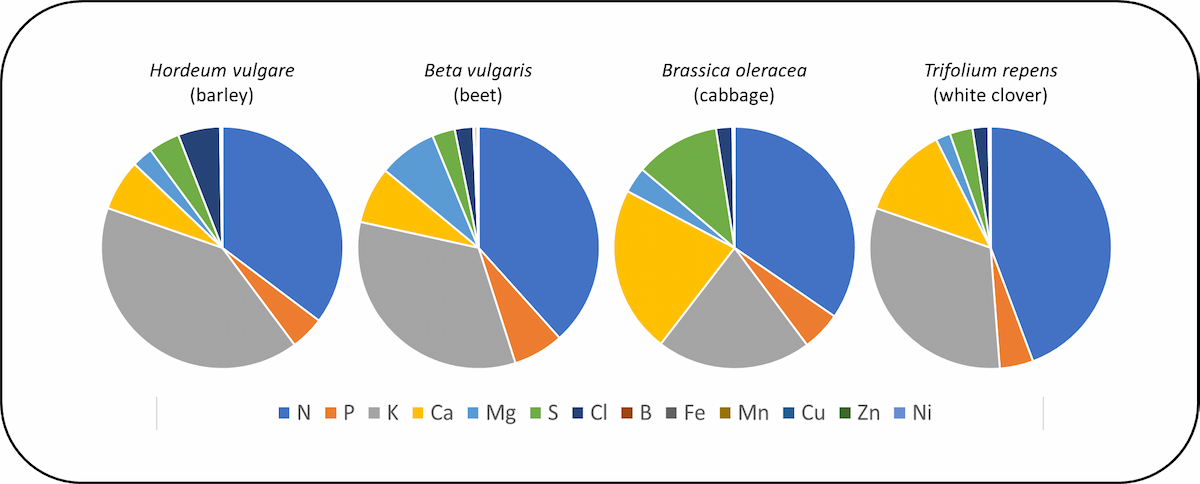
It is clear from recent studies that ionomes differ between plant species (Fig. 4). Detailed knowledge of the distinct ionomes of plant species, and genotypes within plant species, has improved our understanding of the responses of plants to nutrient deficiencies and the development of diagnostics for these, plant ecology and the biogeochemical niche , the genetics of nutrient acquisition and transport within plants, the evolution of plant elemental composition, the choice of species for the phytoremediation of contaminated land , and strategies to improve the mineral nutrition of humans and livestock. Studies of the interactions between elements has provided vital information for a greater understanding of these subjects.
However, can you find these papers? What terms do you search for?
With nutriomics only being used in three or four papers, using this term in a new paper is a bit of a gamble. When other people look for recent research will they remember to include ‘nutriome’ in their search? The papers that will get cited and have an impact on other scientists are the papers that use the same terminology.
If the number of articles published each year on ionomics follows the current trend, then there will be about 55 articles in 2021 and about 75 in 2025. There could be a lot more in 2050, especially if inexpensive instruments for simultaneous, multi-element analysis become available. Thanks to bibliometrics, we can see there’s a future for research into the elemental composition of plants that needs a term. The same bibliometrics also tell us what term most people will be using – ionomics.






