Cataloguing plant diversity and describing new species is a critical and ongoing task that is hampered by a dearth of expertise and an inherently slow process. Even under ideal conditions, collecting a specimen in the wild, describing it as a new species, and publishing that description can take one to two years. More often, it can take decades. Herbaria worldwide are home to a backlog of as many as a million unidentified specimens, and are thought to already contain the majority of undescribed plant species. Computer algorithms taking advantage of machine learning, trained on high quality annotated datasets, could be a key part of the solution.
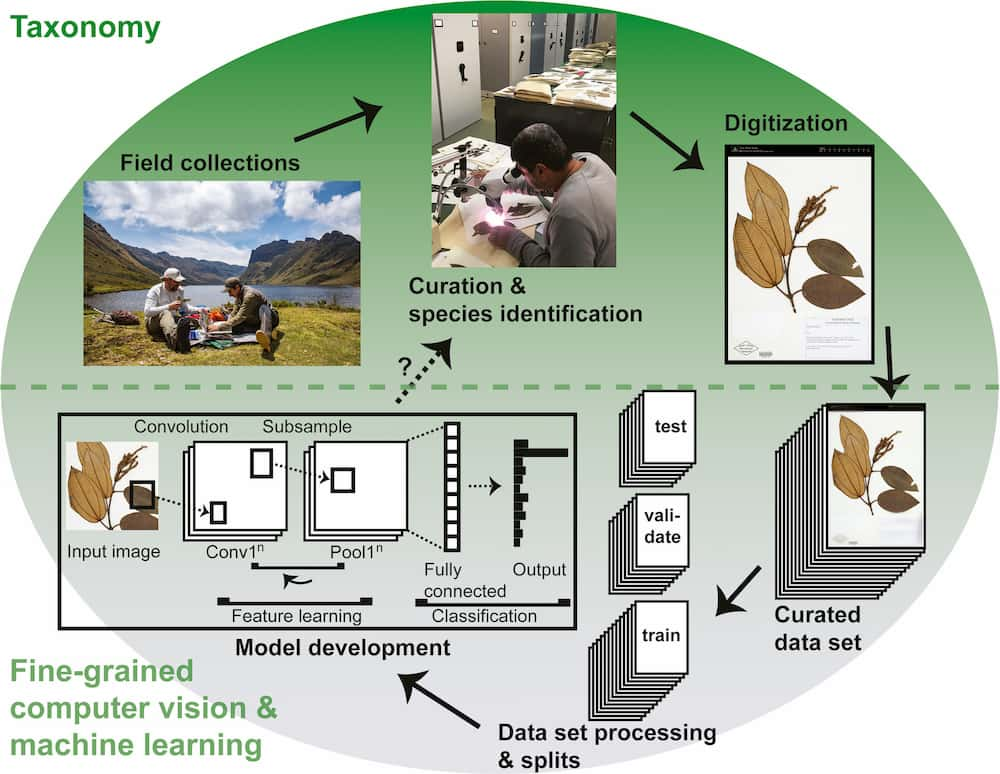
In a new article published in Applications in Plant Sciences’ Machine Learning in Plant Biology special issue, lead author Damon P. Little and colleagues sought ways to harness this potential. The authors hosted a competition on the Kaggle data science platform to develop an automatic species identification algorithm using machine learning. The group put forward a data set for training that included over 46,000 imaged herbarium specimens representing 683 species of the family Melastomataceae. As is typical for herbarium collections, some of these species were represented by many specimens, and others by relatively few.
The competition ran for several months and produced 254 models created by 22 different teams. The four top entries were able to identify species with better than 88% accuracy. The winning teams were from a private company and a public university in China, a team from Facebook AI Research, and, amazingly, one individual, a veterinarian by training, who “joined the competition during his vacation and designed the models on his phone.”
Though the results of the competition were better than expected, only the first step of the problem has been addressed. At present, the algorithms can only assign specimens to the most likely taxon of those it has been trained on; they can’t designate specimens as unknown or new. “The algorithms are not trained ‘to know what they don’t know’, they are trained and constructed to give output based on the training data,” says Barbara Ambrose, a co-author and Associate Curator in Plant Genomics at the New York Botanical Garden. The next step is to formulate an algorithm that can designate a specimen as a probable new species. The authors are currently applying for funding to tackle this challenge.
Ambrose and Little are working to develop a tool that any herbarium can use to work through their unidentified specimens. “The idea is to submit a photo of your specimen and the algorithm will give its top five outputs. We think this will help to clear up backlogs of specimens and can reduce the bottleneck of getting a specimen to an expert in the group. There are many herbaria that are small and may lack taxonomic expertise so this would help them to curate their collections,” says Ambrose. “This is not far off, as Dr. Little has developed a prototype of this that we are calling iCurate. We will need more funding and time to make iCurate more robust in order to benefit herbaria world wide. But hopefully this is not too far off in the future.”
In this vein, Ambrose and Little have recently hosted a second Kaggle competition that vastly expands on the taxonomic scope of the first. “We had 153 teams competing in this competition with a dataset of over 1 million specimens representing more than 32,000 vascular plant species. With the algorithms developed during this competition we are ready to further develop iCurate and tackle the automation of new species recognition.”




