Last semester we started the teaching season with a big challenge: teach an introductory biology class to 320 first-year students. The formal aim of the course was to get the students familiar with plant and animal structures at different scales, and how these structures interact and function together. Aside from that we also wanted to teach them to observe the natural world. We wanted them to realise that they do not need to go to a natural park to see high levels of biodiversity. But how to do that efficiently with 300+ students?
The classical way would be to prepare a deck of slides, detailing the different morphological features of plants and animals, and go through them in a plenary lecture. This is a solution that can be easily upscaled to any number of students. We have been through that as students ourselves. There is no better way to suck the motivation out of a student than reading ten slides describing different leaf arrangements. There had to be a better way. This is why we decided to create BioGO.
BioGO (as in PokemonGO) is a large scale biological treasure hunt we created for our students. The principle is simple. We compiled a list of more than 250 “quests” to be found in Louvain-la-Neuve (the city the Université catholique de Louvain is based in). These quests were all linked to biological structures and organisms, both botanical and zoological. Here are a couple of examples:
- Find an achene
- Find a wood-eating insect
These quests had different difficulty levels (1,2,3) with different rewards. We did not explain the different biological terms to the students in a plenary lecture. We wanted them to look them up and learn by themselves.
Once the students (divided in groups of 3) had a good idea of what was in the list, they had to go outside to find the quests in real life. As we did not want them to damage or collect anything during the game, they had to take pictures of the quests once they found them. We also asked them to activate the geotagging feature of their phones when taking the pictures. These pictures had to be submitted to the student intranet every week, for four weeks. In total, each group had to submit 80 pictures, to reach a sum of 100 points. We would then review them to assess whether they were correct or not.
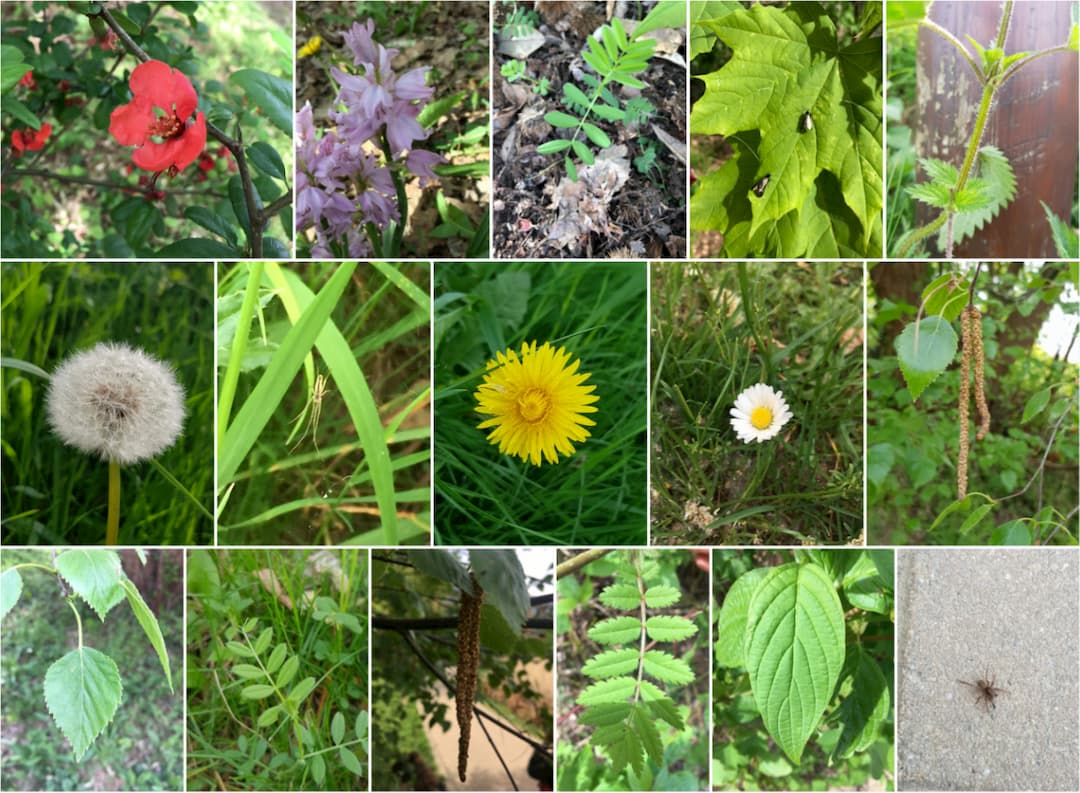

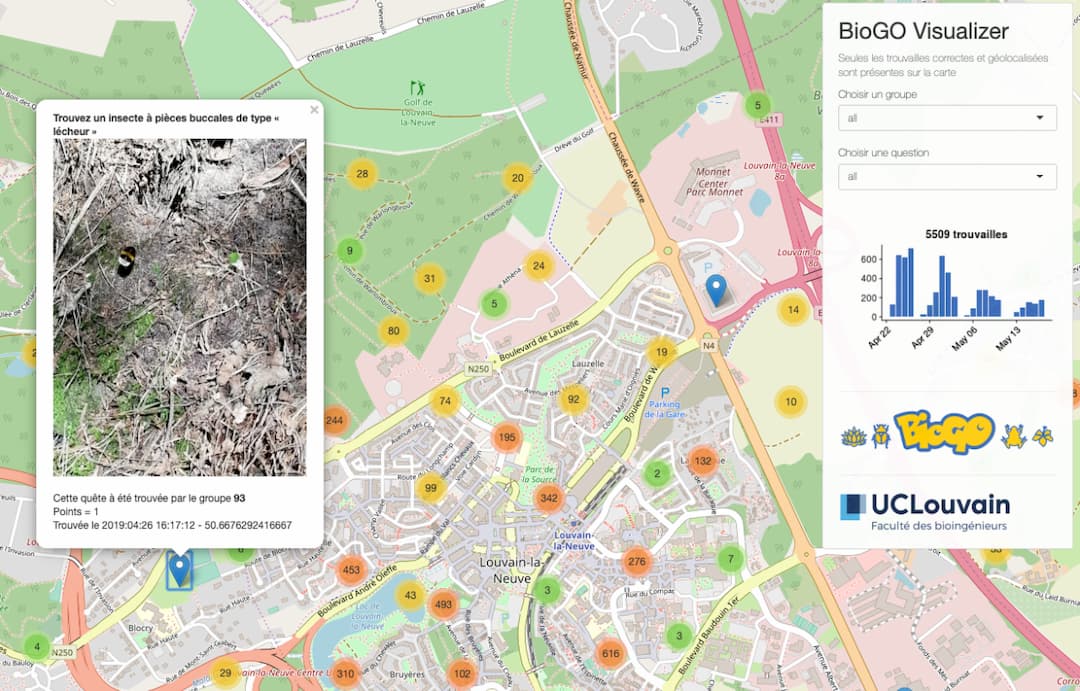
In order to make the experience a bit more fun, and motivate the students even further, we also created a web interface where they could see the number of quests and scores of each team. Each week they would then check their own progress and their place on the scoreboard.
The students collected more than 8000 pictures, more than half of them geotagged and dated. All pictures (and results), for the different quests, can be seen in the final interface of the game (in french): www.biogo.xyz. Our overall feeling was that the students were motivated and eager to find as many quests as possible. We asked them to give us anonymous feedback on their experience. Out of the 211 respondents, 196 (93%) liked the game and 179 (85%) stated that they learned during these four weeks. We also found that, in the following plant biology practicals, students had a better understanding and knowledge of the different botanical terms.
In conclusion, we think the experience was a big success. We achieved a high level of student engagement, and started building a large database of plant and animal biological structures. This database is already valorised in the form of a training web application for botanical terms. We are also currently developing a complete BioGO web interface (free and open-source of course), to allow anyone to use it. We expect that it should be ready to use in the next few months for anyone interested.
About the authors
Guillaume Lobet

Guillaume Lobet is an Assistant Professor, Forschungszentrum Jülich (IBG3, Agrosphere) and the Université catholique de Louvain (Earth and Life Institute). The aim of Guillaume’s research is (i) to understand how various signals that carry information are interacting and being conveyed and integrated at the plant level and (ii) to amplify discrete physiological knowledge into functional plant processes. All of that using Functional Structural Plant Models. More about Guillaume’s research can be found at www.guillaumelobet.be.
Charlotte Descamps

Charlotte Descamps is a PhD student (Earth and Life Institute) and teaching assistant at the Faculty of Bioengineers at UCLouvain. Her time is divided between teaching (mainly botany and plant identification) and pursuing a PhD on plant-pollinator relationships, with Prof. Anne-Laure Jacquemart and Prof. Muriel Quinet. The main objective is to highlight and understand how climate change, through water stress and temperature rise, can affect floral resources and the consequences of these modifications on pollinators.
Lola Leveau

Lola Leveau is a bioengineer, PhD student and teaching assistant at the Faculty of Bioengineers at UCLouvain. The objective of her doctoral research is to compare the performances of innovative cropping systems set up by Belgian farmers, from the agronomic, environmental and economic points of view. This comparison is carried out in collaboration with a network of local farmers, on whose lands the measurements will be taken and with whose advice the protocols and measurement methods have been designed.
Louise Mignard

Louise Mignard is a bioengineer, PhD student and teaching assistant at the Faculty of Bioengineers at UCLouvain. Her thesis’ project deals with the impact of dietary fatty acids on tumour development and progression. The main objective of her work is to highlight and understand, through in vitro and in vivo models, the underlying mechanisms of the strong cytotoxic effects of some unusual fatty acids on advanced-developed tumours which show a high metabolic plasticity and an exacerbated fatty acid metabolism.
Jean-François Rees

Jean-François Rees is an animal physiologist, professor at UCLouvain. He works on fish ecotoxicology at the Louvain Institute of Biomolecular Science and Technology (LIBST). One main aspect of his work deals with the question of the impact of pollutants on deep-sea fish, such as rattails, which cannot be kept alive at the surface, thus forbidding any experimental exposure of the fish to pollutants. For this reason, he develops in vitro systems for investigating the impact of high hydrostatic pressure on deep-sea fish liver cells responses to xenobiotics.





