Though the number of plastome sequences made publicly available has increased rapidly in recent years, intraspecific variation in these sequences is still not well understood in most cases. In Latin America, maize comprises around 260 landraces grown in such differing habitats as the lowlands of southern South America, the highlands of Mexico, and the Andes. Complete chloroplast genomes may vary sufficiently at the sequence level to help distinguish the evolutionary trajectories of developing landraces over time.
In a recent article published in Annals of Botany, lead authors Mariana Gabriela López and Mónica Fass and colleagues attempted to estimate the degree of plastome diversity within Latin American maize by studying various landraces as well as several wild teosinte relatives. The group sequenced the complete plastomes of 30 landraces and three teosintes, using these sequences in combination with those publicly available. The data was used to quantify haplotypes and infer evolutionary relationships among them.
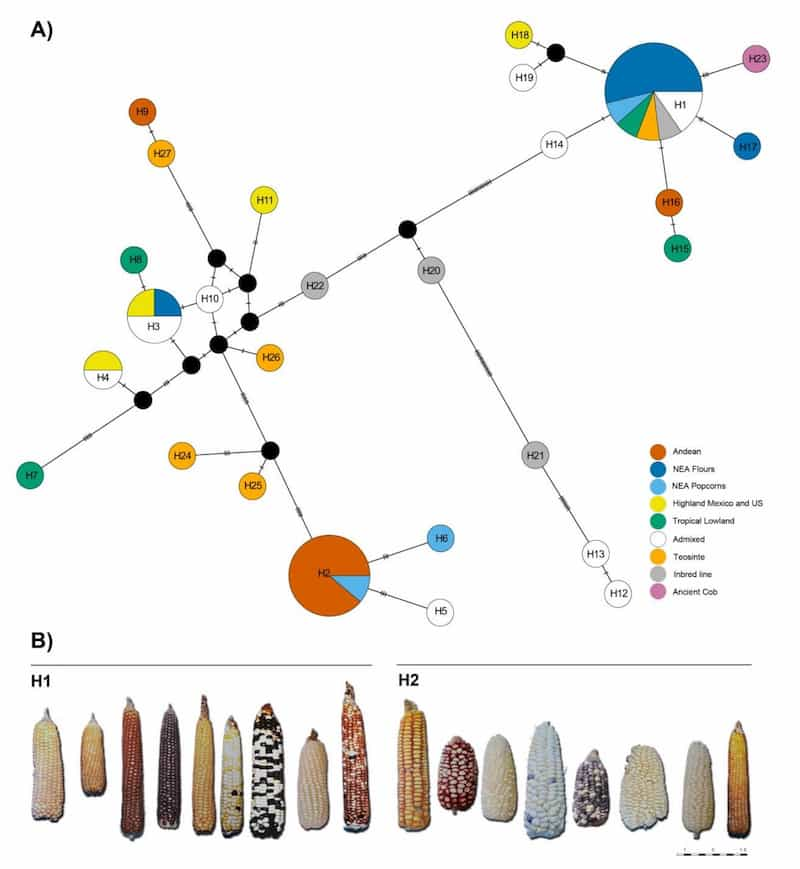
Sequencing produced 124 polymorphic plastome loci circumscribing 27 different haplotypes across 51 maize individuals. Significantly different haplotype distribution could be seen in Andean versus lowland landraces, a result “in accordance with the differentiation of Andean and lowland lineages from a secondary South American domestication center,” write the authors. “However, our results are also compatible with the existence of two independent diffusion waves accounting for the diversity of South American landraces,” they add.
Surprisingly, overall, maize and teosinte plastomes showed a scattered distribution that made them uninformative in terms of tracking sub-species diversification. Furthermore, fully 85% of the uncovered haplotypes were present in only a single individual, hinting at high levels of underlying genetic diversity. “[G]iven that plastid markers are the tools of choice for most plant systematics studies, knowledge of intraspecific variation may allow for a more comprehensive understanding of evolutionary processes at low taxonomic levels, particularly when species delimitation is unclear,” write the authors.





