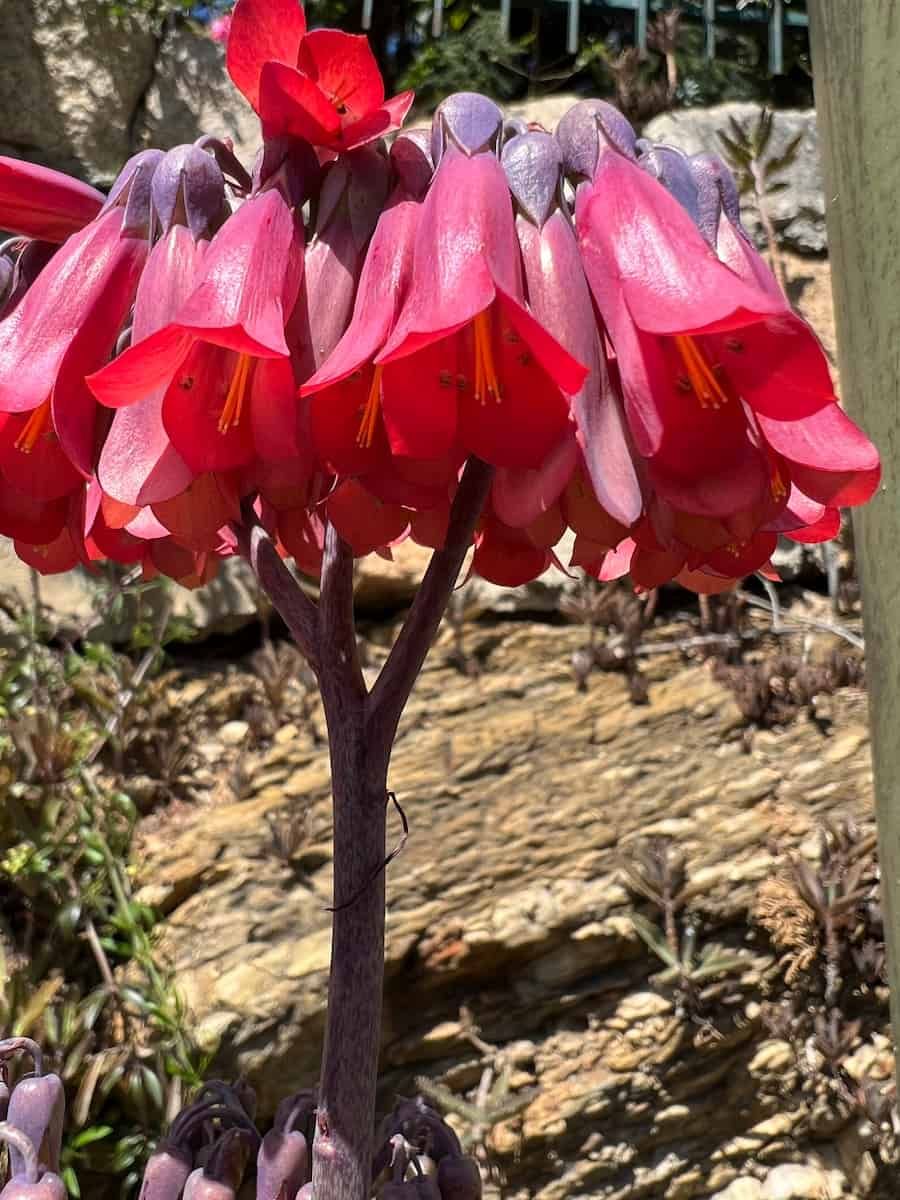
Recent work on the green alga Haematococcus lacustris uncovered the largest plastid genome on record: a whopping 1.35 Mb (with >90% non-coding DNA). In a recent review published in AoBP, David R. Smith takes a closer look at this gigantic plastome, comparing it to other large organelle DNAs. David first came across the H. lacustris chloroplast genome while scanning the newest cohort of plastomes and, to his dismay, found that it had been written up in a non-peer reviewed journal that rapidly publishes short reports on new microbial chromosome sequences. He decided that the H. lacustris plastome deserved to be noticed and appreciated properly by those in the chloroplast research community, and a review was born.
In his review, David shows that the H. lacustris plastid coding repertoire is not as unusual as initially thought, representing a standard set of rRNAs, tRNAs, and protein-coding genes. The intergenic spacers are dense with repeats, and it is within these regions where potential answers to the source of such extreme genomic expansion lie. By comparing two closely related strains of H. lacustris, David argues that the mutation rate of the noncoding plastid DNA is high and contributing to plastome inflation. With the growing importance of H. lacustris as an industrial alga, there will likely be a lot of sequencing data arriving to GenBank in the coming months and years— music to David’s ears as this unicell with its behemoth of a plastome surely has a lot more to teach us about organelle genome evolution.
Researcher highlight

David completed his BSc at Acadia University in beautiful Wolfville, Nova Scotia, Canada in 2005, and then moved 75 km down the road to Halifax for a PhD in Genetics at Dalhousie University (2005–2010), exploring the weird and wonky genomes of green algae. Tired of the Atlantic Ocean, he travelled across the country to Vancouver for a postdoc in the Biodiversity Research Centre at the University of British Columbia (2011–2013), continuing to study genome evolution in algae. He started an assistant professorship in the Biology Department at the University of Western Ontario in 2013 and was promoted to Associate Professor in 2018. His group spends too much time pondering the origins of non-photosynthetic algae (evolutionary burnouts). They also do a lot of science writing and science communication work. You can find them online at http://www.arrogantgenome.com. Outside the lab, David likes to run marathons and drink wine, but rarely at the same time.



