Climate change will alter the availability of resources affecting plant performance. One way plants will respond to these changes is through shifts in phenotype (i.e. traits). The ability of individual genotypes to produce different phenotypes when exposed to unique environmental conditions is called phenotypic plasticity. Understanding plastic responses is crucial for predicting and managing the effects of climate change on plants. Computational plant modelling offers insight into plant–environment interactions and resulting phenotypic plasticity.
Dr. Romain Barillot and colleagues, researchers at the French National Institute for Agriculture, Food, and Environment (INRAE), used a computer model to explore the processes underlying shoot morphogenesis – the biological process that causes shoots to develop their shape.
Typically, wheat produces many branches (i.e. culms) but plants lacking the gene(s) that control branching present a monoculm phenotype. The use of monoculms allows researchers to focus on simulating leaf growth plasticity without needing to consider the putative rules of tiller emergence. For this reason, the authors adapted CN-Wheat, a mechanistic model which fully integrates shoot morphogenesis and the metabolism of carbon (C) and nitrogen (N) at organ scale, within a three-dimensional representation of plant architecture, to simulate the growth of monoculms.
According to Barillot and colleagues, “leaf plasticity in previous grass models is defined from empirical data, therefore limiting our ability to explore the response of plants to the wide range of novel growth conditions predicted due to climate change. What is unique about CN-Wheat is that it explicitly considers the role of carbon and nitrogen economy of the whole plant in interaction with the environmental conditions. The model uses feedback loops between carbon and nitrogen acquisition and responses to metabolite concentrations to drive morphogenesis, allowing us to mechanistically explore the response of plants growing under a wide range of environments.”
The authors used the modified CN-Wheat to explore how morphogenesis under a variety of growth conditions affected the plasticity of major foliar traits. Simulations were run with plants growing under a range of interacting planting densities, soil N concentrations, and incident photosynthetically active radiation, which is the light available for photosynthesis. The authors then assess the plant’s dimensions, biomass, specific leaf area, and nitrogen content.
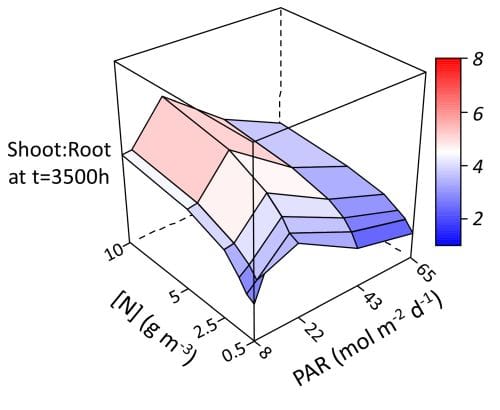
The authors found that the model simulated realistic phenotypic plasticity for contrasting light availability and N fertilization. For example, as a response to increasing incident PAR, the model simulated, at the whole-plant level, a decreasing fraction of dry mass allocated to roots (Fig. 1) and faster leaf emergence, and, at leaf level, thicker and wider leaves (Fig 2). Also, as a response to increasing soil N concentration, the model simulated, at the whole-plant level, a decreasing shoot:root dry mass (Fig. 1) and faster leaf emergence, and, at leaf level, increasing thickness, width and length (Fig 2).
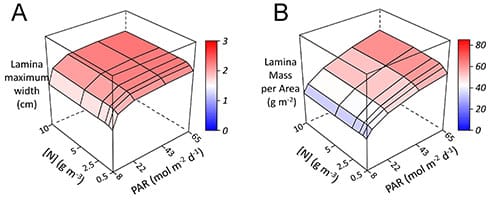
These results demonstrate that integrating plant functioning at organ scale can simulate, as an emergent property, the phenotypic plasticity of plants in contrasting light and nitrogen conditions.
Barillot concludes, “CN-Wheat provides an original and explicit description of the processes underlying morphogenesis, which offers new possibilities to link the phenotypic plasticity of plants to their C and N metabolic status at the place and time where traits are built. This unique level of integration makes CN-Wheat a good tool to explore plant functioning in contrasting environments and opens up new possibilities to account for genotype-environment interactions.”
READ THE ARTICLE:
Marion Gauthier, Romain Barillot, Bruno Andrieu, Simulating grass phenotypic plasticity as an emergent property of growth zone responses to carbon and nitrogen metabolites, in silico Plants, 2021, diab034, https://doi.org/10.1093/insilicoplants/diab034
This manuscript is part of in silico Plant’s Functional Structural Plant Model special issue.






