Linking genome sequence mechanistically to complex plant phenotypes requires significant biochemical and biophysical detail. Unfortunately, plant gene regulatory network models represent their RNA components with arbitrary mass units. This practice is inherited from the experimental methodology, where RNA abundance is typically normalized to an internal standard for quantification, yielding data in arbitrary, relative units.
The use of arbitrary units prevents researchers from assessing the validity of the values and do not provide the biochemical and biophysical detail required for biological engineering.
Uriel Urquiza García, Postdoctoral Researcher at Heinrich-Heine-Universität Düsseldorf, and Andrew Millar, professor at The University of Edinburgh, improve on an existing plant gene regulatory network model of the clock gene circuit by refactoring the model to use absolute mass units in their new in silico Plants article.
The authors focused their efforts on a mathematical clock model. “Plant clocks coordinate many developmental and physiological processes. Farmers and breeders have extended the range of cereal crops northwards, unaware that they were selecting for mutations in clock- and clock-related genes. Understanding the clock gene circuit promises to guide further breeding and/or engineering of crop varieties, for new combinations of climatic conditions,” says Millar.
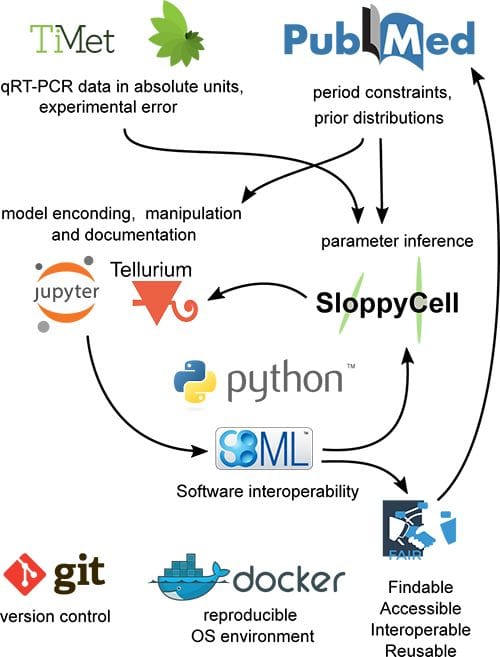
The authors re-factored an existing clock model, P2011, to use absolute values of clock gene mRNA levels in units of RNA transcripts per cell, which were published by their earlier project “TiMet” (see figure). The resulting alternative model was termed U2019. The U2019 model was further updated by including additional circadian regulatory interactions to produce the model U2020.
According to Millar, “by recalibrating our models into absolute units of transcripts per cell, it became possible to test the modelled transcription rates against measured biochemical data. Unfortunately, we had no “ground truth” data for plant transcription rates in general. Uriel combined two datasets from the literature to create the reference dataset of measured plant transcription rates that we needed.” The authors found a highly significant correlation between the published data sets, which allowed the data to be combined for this purpose.
Testing the models’ inferred transcription rates compared to the reference dataset represents an advance in biochemical realism for models of plant gene regulation.
RESEARCH ARTICLE
Uriel Urquiza-García, Andrew J Millar, Testing the inferred transcription rates of a dynamic, gene network model in absolute units, in silico Plants, Volume 3, Issue 2, 2021, diab022, https://doi.org/10.1093/insilicoplants/diab022
The authors have taken an Open Science approach, so their reference transcription rate distribution can be unambiguously referred to using its identifier in the BioNumbers database, BNID117324. The data, models, computational environment as a Docker container, and other resources are publicly available from the FAIRDOMHub.org repository, as a static Snapshot, structured according to the standard ISA hierarchy (doi: 10.15490/FAIRDOMHUB.1.INVESTIGATION.170.3). See the article for additional details.






