Circadian clocks in plants produce an internal estimate of time that synchronizes biological events with day and night. They play a critical role in plant physiology, growth, development, and survival. Therefore, the inclusion of circadian rhythms in crop growth models is essential for high fidelity results.
While there are existing circadian clock models, they are often inflexible or result in complex outputs, limiting their use in general simulations.
Postdoctoral Researcher Edward Lochocki and USDA-ARS Research Plant Physiologist Justin McGrath, both of University of Illinois, developed a new circadian clock model that easily adapts to different environmental conditions and produces intuitive outputs.
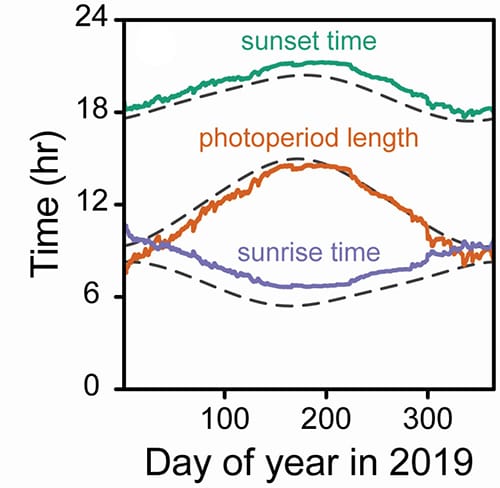
Real-world clocks use physical oscillators such as pendulums to keep time, and the authors used the same approach to model circadian clocks with mathematical oscillators. The model included two oscillators that were controlled by the onset of dawn and dusk, which were able to account for the change in daylength over a year.
“Many crop models calculate daily sunrise times and photoperiod lengths using celestial mechanics, which are mathematical representations of solar coordinates that are used to calculate sunrise, sunset, and photoperiod length,” says McGrath. “While this approach is readily understood and straightforwardly used as inputs to other model components, it does not provide an intuitive link between genetic components and plant behavior. Our oscillator model has components easily related to known genetic components of the circadian clock. This allows easier incorporation of environmental influences, such as temperature, by altering the kinetics of the modeled oscillators.”
The authors compared the predicted flowering dates for soybean sown throughout the year by the CROPGRO soybean development model using either their oscillator clock model or celestial mechanics. They found that flowering time using the oscillator clock model closely matched the performance of celestial mechanics.
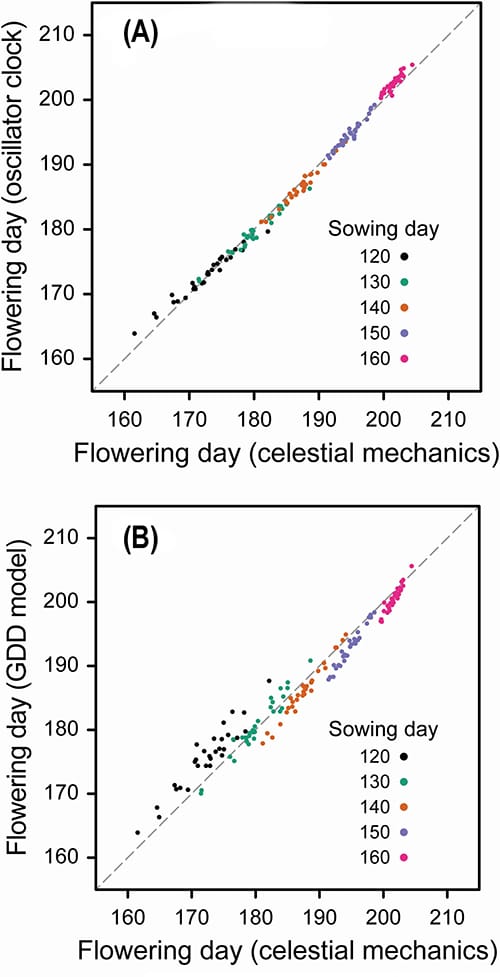
The authors also compared the predicted flowering dates for soybean sown throughout the year by running CROPGRO with either their oscillator clock model or growing degree days (GDD). GDDs use accumulated days with higher than minimum temperature to predict life cycle transitions in plants. They depend solely on temperature and are used to estimate soybean development, where developmental stages are assumed to occur at particular GDD thresholds.
According to Lochocki, “Although modeling the molecular components of the circadian clock provides the most intuitive way to examine how genetic alterations affect plant behavior, such a model is too computationally expensive to solve as part of a full crop growth simulation for an entire season. A simplified model that captures the overall functionality of the gene network allows one to study the broad behavior of the system, if not the details, in a mathematically tractable way.”
Another approach to modeling the circadian clock is the gene network clock, which are systems of differential equations representing gene networks that function independently of the light source. While they link genetics and plant behavior, they require detailed knowledge of the regulatory pathway between the clock and the relevant genes to be accurate.





