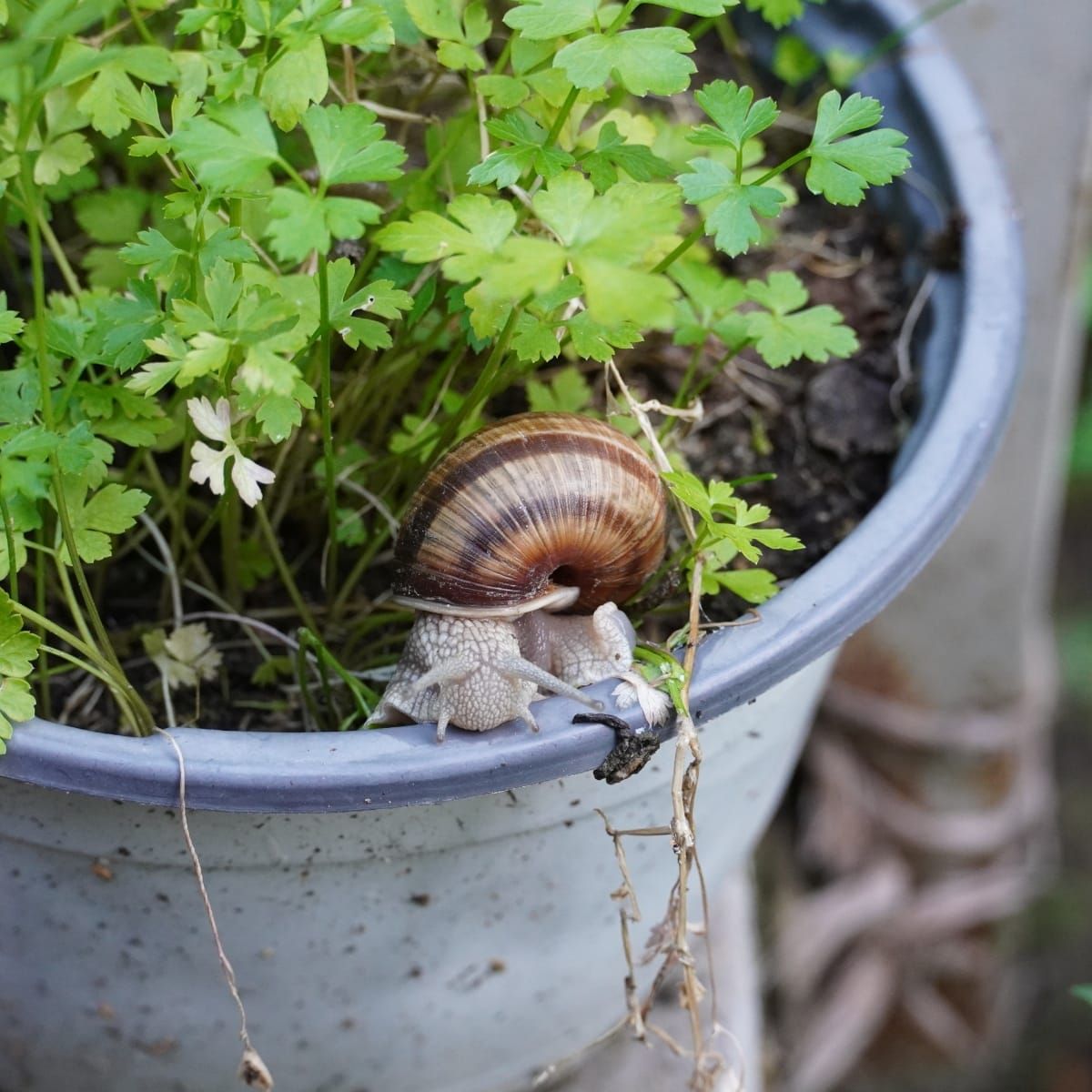
Soybean is one of the largest sources of vegetable oil and of animal protein feed in the world with an estimated global export value of $58 billion USD in 2018.
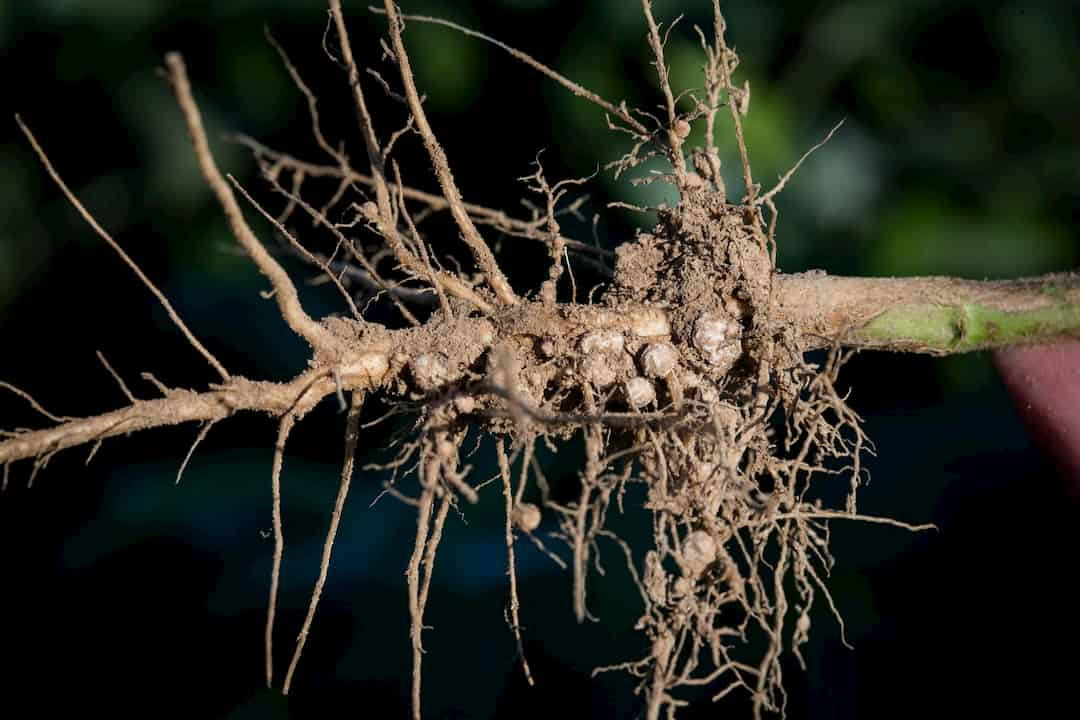

Large amounts of nitrogen are needed for soybeans due to their high seed protein content, and soybean obtains 50-60% of it from the atmosphere. This is possible due to symbiotic interactions with nitrogen-fixing bacteria collectively called rhizobia, which pull nitrogen from the atmosphere and convert it to a form the plants can use. This process reduces the need for chemical nitrogen fertilizers, which are expensive and cause environmental pollution.
Rhizobia are only present within root nodules, which are a root lateral organ. Understanding the regulation of genes involved in lateral root and nodule development will inform biotechnological strategies to optimize nodule formation and enhance nitrogen fixation.
In a paper recently published in in silico Plants, Dr. Shuchi Smita and coauthors present a robust computational framework to predict potential gene regulatory networks (GRNs) associated with root lateral and nodule development in soybean. This framework integrated genomic and transcriptomic data to infer the key regulators and GRNs associated with nodule development in soybean.
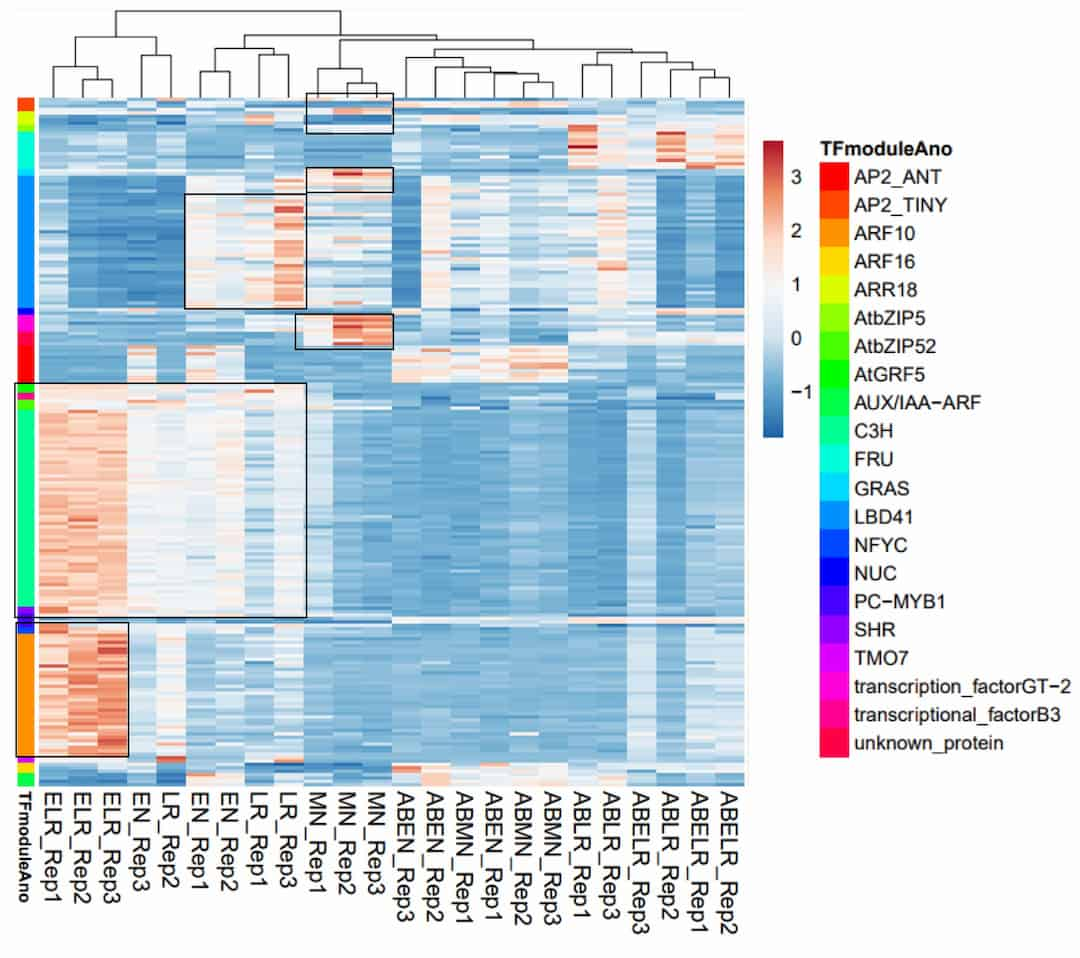
This study identified 21 gene regulatory modules of 182 co-regulated genes networks potentially involved in root lateral organ development stages in soybean, including previously known nodule- and lateral root-associated transcription factors.
Senthil Subramanian, Associate Professor of Plant Science at South Dakota State University, explains “understanding molecular mechanisms behind genotype to phenotype relationships will help determine key gene targets for genetic or biotechnological strategies that can optimize nodule formation and enhance nitrogen output.”





