Green or purple, round or elongated, soft or hard, stuffed or not. Choosing table olives and olive oils can become an overwhelming experience because of their variety and tastiness. The domestication of Meditterrenean olive trees (Olea europaea subsp. europaea) is estimated to date back 6,000 years in the Levant region. There are many questions about olive domestication. Is there a diversity hotspot of cultivated and wild olives (O. europaea subsp. sylvestris)? How were the different cultivars selected, where and why?
Drs Irene Julca and Toni Gabaldón with eight colleagues from Barcelona Supercomputing Centre, The Barcelona Institute of Science and Technology University of California Irvine, University of Cordoba, Universitat Pompeu Fabra and Royal Botanical Garden of Madrid, assembled the whole genomes of 46 varieties (O. europaea subsp. europaea) and 10 wild olives (O. europaea subsp. sylvestris) to detangle their domestication processes. The researchers found that overall, cultivated olives have only slightly lower levels of genetic diversity than wild forms and cultivar selection has not preferred varieties that carry specific genes responsible for fruit size or oil content but rather changes in gene expressions.
This group of scientists assembled the first olive tree genome (Oe6) in 2016 but lots of questions emerged after the genome publication of a wild form from Turkey in 2017. As the pollen of olive trees can be carried far away by wind and cultivated trees can cross-fertilise with wild olives (O. europaea subsp. sylvestris), it is hard to tell which tree is cultivated or a wild form. The World Catalogue of Olive Varieties from 2000 describes 139 varieties and shows the different phenotypes of olive trees.
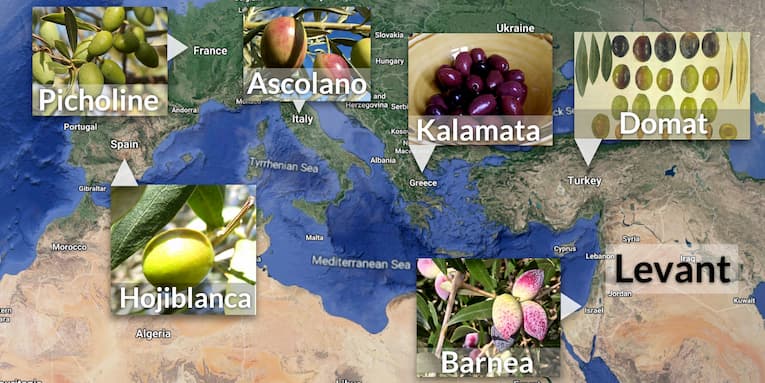
Julca and colleagues selected authenticated cultivars from the olive germplasm collection at Cordoba University and sampled wild forms from a coastal area of Spain where there were no historically cultivated olive tree plantations. The researchers re-assembled the previous genome (Oe6) of cultivar (cv.) Farga and could characterise an additional 4,911 genes using the new genome (Oe9) assembly. They also assembled the plastid, mitochondrial and nuclear genomes that are inherited in a different way (e.g. organelles are maternally inherited) of 46 cultivars and 10 wild forms.
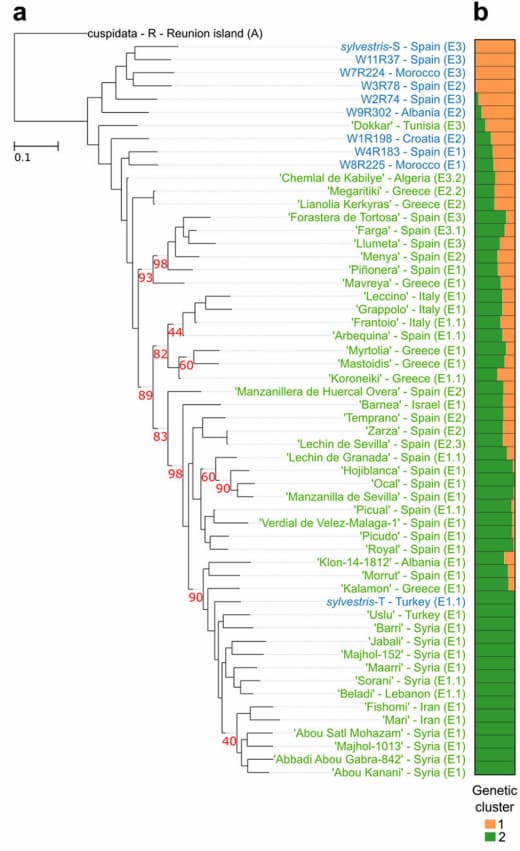
The researchers found that over 1,000 genes were absent from the wild form genome compared to the genomes of cultivars. These genes were mostly associated with stress response, growth and development. Whilst it is common that cultivated fruits go through a bottleneck event (e.g. human selection) and have lower genetic diversity, it was not the case for olives. Genetic diversity was only slightly smaller amongs cultivars compared to wild forms. Whilst fatty acid metabolism and accumulation was thought to be one of the most important characters in olive domestication, Julca and colleagues could not find any signs of selection of these genes. The only potential genes under selection related to changes in gene expressions.
Phylogenetic analyses estimated a mild population bottleneck 3,000-14,000 years ago but afterwards population size expanded and some genes moved from wild forms to the cultivated varieties multiple times. Nuclear genomes suggested that that most cultivars mainly derive from a common primary domestication process whilst the plastid genomes suggested genetic contributions from three different genetic pools. A genetic structure analysis suggested that there were two distinct ancestral genetic pools which are differentially present in the 56 individual plants, forming three clusters (e.g. one containing more of wild forms and two containing different levels of mixtures of the two genetic pools). Cluster 3 mainly consisted of eastern cultivars from Syria, Iran, Lebanon and Turkey.

“[T]his study represents the largest phylogenetic analysis of genome-wide sequences of the Mediterranean olives,” Janca and colleagues wrote.
The researchers suggest that olives represent a domestication continuum where there was a primary domestication event followed by many hybridisations amongst cultivars and wild forms.
“Our results, together with those from previous analyses, suggest that cultivated individuals have similar nucleotide diversity as compared with wild individuals, being slightly higher in cultivars admixed with the western wild genetic pool, possibly due to introgression with local wild populations,” the researchers wrote.
In a related article, Dr Toni Gabaldón wrote, ”One important gap in the existing data is the lack of representatives in the whole genome datasets from the wild genetic pool of the Eastern Mediterranean, the original genetic pool from which the first cultivated olives were likely selected”.
The genome assemblies and improved reference genome pave the way to more exciting research about olives. They also provide an interesting lesson about crop domestication. Some crops have been “overseleted” for specific traits (and the genes responsible for them) that decreased genetic diversity between cultivars and now, scientists are trying to find wild relatives to introduce some diversity and other traits. There are many plant diseases and pests threatening olives and their genetic diversity will be important to find resistant varieties.
The most olive oil and table olives are consumed in Greece (9.4 kg/year/per capita) and in Cyprus (3 kg/year/per capita) within the EU according to the International Olive Council. Olives are not only important in a culinary sense but the trees also have climate change mitigation functions.
“We hope future collaborative efforts will help us increase our dataset with relevant and representative varieties from wild and cultivated olives, we would be happy to collaborate in this regard”, Gabaldón wrote.





