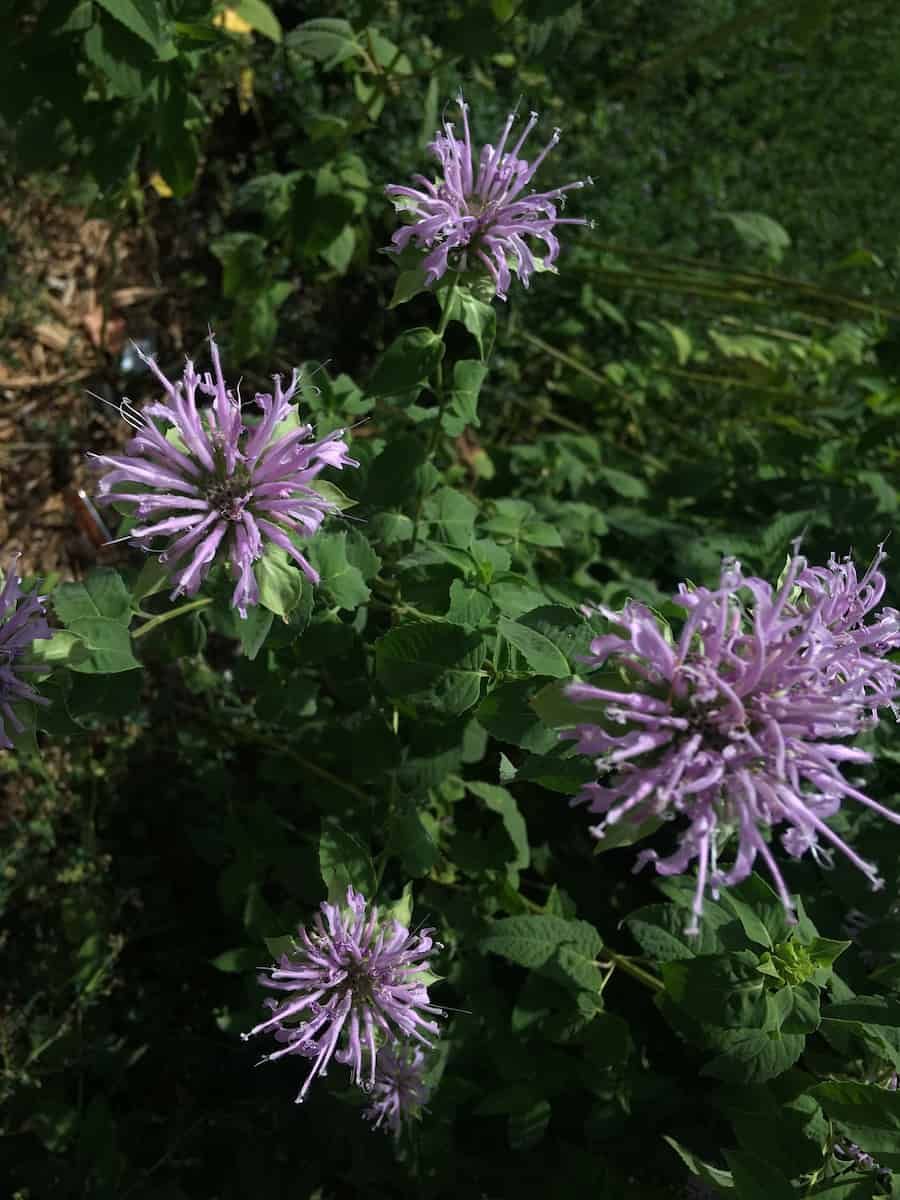
With new technology and techniques leading to new specimen uses and means for analysis, natural history collections are only becoming more valuable as time goes by. Limitations in sequencing technology had until recently made sequencing very old herbarium specimens next to impossible. Advances in next generation sequencing (NGS) are continually pushing back the age at which samples are still viable, opening up avenues for the analysis of type specimens and recently extinct taxa that are over a century and a half old. This work has the potential to refine the systematics of numerous groups.
In a new article published in the Botanical Journal of the Linnean Society, lead author Laurent Hardion of the University of Strasbourg and colleagues used NGS to sequence the whole plastid genome of a 167-year-old herbarium specimen of Leptagrostis schimperiana. This grass species, last seen alive over 150 years ago, is one of two remaining incertae sedis, or members of unknown placement, of the Poaceae subfamily Arundinoideae, once a dustbin group that has been steadily resolved over the last few decades. The other, Piptophyllum welwitschii, was analysed morphologically along with L. schimperiana to determine its proper phylogenetic placement.
Analyses showed that L. schimperiana belongs in the Arundinoideae tribe Crinipedeae, the largest of the three tribes in the subfamily, with distribution across southern and eastern tropical Africa. Though P. welwitschii was not successfully sequenced, morphological analyses also placed it within Crinipedeae, which is relatively homogeneous morphologically, and circumscribed by such specific characters as caepitose plants; solid culms; small, few-veined glumes; and bifid or winged paleas.
The success of sequencing ventures such as this is part of a trend of increasingly ambitious genetic work on old specimens, but there is much to do. “Herbarium collections contain numerous specimens with taxonomic ambiguities,” says Hardion. “It is a major problem, because the most important specimens for species description are often old specimens, and previous sequencing methods did not allow us to study them. Thus, this next generation sequencing method will resolve numerous questions in biodiversity description based on herbarium specimens. The residual problem is the partial impact on the specimen, because we need a piece of plant material for DNA extraction, and specimen sampling is sometimes not possible on small and very precious specimens.”
Hardion notes that this technique is “still underused” on old specimens, and may work on even older material than what his group used. He knows of an as-yet unpublished case of successful plastid genome sequencing of a voucher collected during the eighteenth century. “The most important question is of the preservation conditions of the sample since its collection,” he says.




